
Bonus:¥500000
118
Teams
340
Registrations
Deadline:2025-07-12 23:59:59
My team
Team LeaderTo be established
In review
Established
Deadline:2025-07-12 23:59:59
Upload Works
To be uploaded
Uploaded
Deadline:2025-07-12 23:59:59
Event Introduction
Academic Committee
Competition Task
Awards and Benefits
Cloud Resources Access
I. Background
In recent years, countries worldwide have attributed high importance to the development of computational biology, leveraging its methods and technologies to address challenges in the biopharmaceutical industry. To further advance this field, "Shanghai International Computational Biology Challenge" will be held consecutively to engage universities, research institutions, enterprises, and individuals globally. The competition seeks to identify outstanding talents and projects, build a talent and project pool, and strive to become an influential international event in AI-driven life sciences.
To further advance computational biology and implement the Shanghai Computational Biology Innovation Development Action Plan (2023-2025), the Shanghai International Computational Biology Challenge 2025 will be held consecutively to engage universities, research institutions, enterprises, and individuals globally. The competition seeks to identify outstanding talents and projects, building a talent and project pool, and strives to become an influential international event in AI-driven life sciences.
"Shanghai International Computational Biology Challenge 2025" focuses on HCAR1, utilizing its known protein sequence and structural information to screen potential drug candidates through AI computation—via both small molecules (lactate receptor antagonists) and large molecules (lactate receptor-specific antibodies). Wet-lab experiments will validate their specificity and activity, accelerating HCAR1-targeted drug discovery and clinical translation while promting the practical application of AI in drug development.
II. Objectives
To establish an end-to-end innovation system encompassing technology innovation – drug discovery – translational application, leveraging Shanghai's dual advantages in clinical resources and computational biology to foster internationally competitive original drug targets and inject new momentum into scientific innovation and industrial upgrading.
To attract top global talent in computational biology, strengthen interdisciplinary collaboration, exchange cutting-edge innovations, explore real-world demands, and spark creative breakthroughs to enhance Shanghai's overall competitiveness in computational biology.
To develop the "Shanghai International Computational Biology Challenge 2025" into a globally recognized brand, elevating Shanghai's international influence in computational biology.
To foster an ecosystem for innovation and application, focusing on frontier technologies and practical solutions. And to implement a competition-based evaluation and talent selection mechanism to attract more innovation, industry, and capital resources, forming a resource network to advance the commercialization of original algorithms, software, and structural technologies.
III. Participants and Requirements
(1) Eligibility: Open globally to individuals and teams from higher education institutions, research institutions, enterprises, and other organizations, regardless of age or nationality. Participants may register and participate via the official website(https://www.ibiws.com/2025/SICBC?lan=en).
(2) Registration Requirements: All team members must complete registration and team formation on the official website. Participation is confirmed upon successful real-name verification.
(3) Team Formation Requirements:
-Each participant may only create or join one team. Changing teams is not permitted.
-Each Team must consist of 1 to 5 members.
-The team leader must create the team using the〈My Team〉section on the competition official website (the team name is chosen by the leader).. Other team members must search for and join the team using its name.
- All participants must complete team formation before the registration deadline.
※Participating teams may choose to compete in the small molecule track (lactate receptor antagonists), the large molecule track (lactate receptor specific antibodies), or both tracks.
Additional Rules:
1. Individuals or organizations involved in competition design are ineligible.
2. Accounts are personal and non-transferable.
3. Participants must ensure information accuracy; misuse or falsification will result in disqualification.
4. Account security is the participant's responsibility; reaches must be reported immediately.
IV. Competition Topics and Data
1. Topic Overview:
On February 4, 2025, a research team from East China Normal University published in Nature Immunology, revealing that HCAR1 (a GPCR family member) drives tumor immune evasion by sensing lactate signals. The study highlighted HCAR1 inhibitors’potential in cancer immunotherapy.
HCAR1 is a promising anti-tumor drug target, but high-affinity, selective drug molecules remain undeveloped due to challenges like high-throughput screening difficulties and structural data scarcity.
As a prototypical cell-surface receptor, HCAR1 can specifically recognize lactate molecules and mediates downstream signaling pathways. It plays a central role in regulating critical physiological processes such as energy metabolism and immune-inflammatory responses. Latest studies have shown that overactivation of HCAR1 signaling is closely associated with various pathological processes, including metabolic disorders and tumor progression. For instance, HCAR1 overactivation can enhance the recruitment of immunosuppressive cells, thereby inhibiting anti-tumor immune responses and driving malignant progression in cancers such as colorectal cancer. Additionally, abnormal activation of HCAR1 has been implicated in tumor-induced cachexia by promoting heightened metabolic activity in adipocytes.
At present, although the therapeutic potential of HCAR1 as a drug target is increasingly recognized, there remains a significant gap in the development of high-affinity, highly selective HCAR1-targeting drug molecules. The major obstacles hindering progress in this area include the technical challenges of high-throughput activity screening, the lack of detailed three-dimensional structural information, and interference from multiple homologous receptors.
The competition will employ structure-based AI screening (antagonists/antibodies) with experimental validation to expedite HCAR1 drug development and showcase AI's pharmaceutical applications
(For detailed competition topics, please refer to the Competition Details page.)
2. Small-Molecule Compound Library:
(1) The library comprises approximately 10 million compounds along with detailed information (a download link to the compound library will be provided: Computational Compound Library.zip). It includes around 20,000 compounds with known bioactivity, and approximately 10 million additional compounds drawn from a curated commercial selection and an extended compound collection.
(Library construction criteria: (1) Deduplication based on structure standardization and hash indexing methods; (2) Removal of unrecognizable molecules using the RDKIT algorithm; (3) Retention of compounds with molecular weights between 300 and 600 Da; and (4) Software compatibility testing.)
(2) To encourage the discovery of original new drugs, this competition accepts de novo generated novel compound samples submitted by participating teams.
3. Antibody Sequence Library:
- INDI database (http://research.naturalantibody.com). Participating teams may perform high-throughput screening of existing databases to identify HCAR1-binding antibody candidates, or further engineer/modify them.
- Only sequences (IMGT-numbered) are required. The organizing committee will be responsible for all antibody expression and activity validation. To ensure standardization, all antibodies must use the IMGT numbering scheme.
V. Competition Process
The competition consists of three phases: Registration, Preliminary Stage, and Finals Stage. Small molecule (lactate receptor antagonist) and large molecule (lactate receptor-specific antibody) tracks will be evaluated separately, following the same competition timelines:
(A) Small molecule (lactate receptor antagonist) Competition Schedule:
1. Registration(From now - July 12 ,2025, 23:59:59 UTC/GMT+8)
Participants may register as individuals or in teams (all entrants must form a registered team). All required submissions must be completed within the specified timeframe. The organizing committee will announce the selected teams and the list of small molecule compound by July 25, 2025.
Required submissions from participating teams include:
Time | What to Submit | Requirements |
From now - July 12 ,2025, 23:59:59 UTC/GMT+8 | Proposal abstract | Participating teams must submit a proposed summary for their planned research using the official template (Attachment 1. Template for Proposed Project Abstract.docx). The Organizing Committee will select maximum of 100 teams based on research competency (prior work in relevant fields), problem comprehension (understanding of HCAR1 challenges), implementation feasibility (methodological soundness) and innovation potential (novelty of approach). Please find detailed scoring rubric in Attachment 2. Abstract Scoring Criteria.pdf. |
Candidate molecules | Participating teams will compete using self-developed AI computational models, submitting a list of 100 molecules ranked by predicted score from highest to lowest. Based on this submitted prioritization, the Organizing Committee will select a minimum of 10 molecules per team for wet-lab validation. Please download template (Attachment 3. Preliminary Round Submission Template.docx) | |
The source code of the algorithm capable of reproducing molecule screening results | The Organizing Committee strongly encourages all participating teams to submit reproducible source code for molecular screening, promote open-source licensing and advocates for original algorithm development. Teams selected to advance in the competition must submit their source code. |
2. Preliminary (July 25 –early October, 2025)
The organizers will conduct wet-lab validation and evaluation of submitted molecules (Attachment 4. Experimental Testing Procedure and Scoring Criteria for the Small Molecule Preliminary Round.pdf).
The Organizing Committee will announce the teams advancing to the Finals in mid-October 2025.
The Preliminary Round consists of the following three phases:
Time | Phase | Schedule |
July 25 –early October, 2025 | Screening | Based on dual-concentration antagonistic activity evaluation using GloSensor cAMP assay, the top 100 active molecules(ranked by antagonistic activity) will advance to the next phase. |
Evaluation | Following multi-concentration antagonistic activity evaluation and preliminary comparison using GloSensor cAMP assay, the top 10 ranked teams will advance. | |
Review | ·Stage 1: After multi-concentration antagonistic activity evaluation using HTRF cAMP assay, the top 5 ranked teams advance to Stage 2(and simultaneously qualify for the Finals). Teams entering the Review phase but not advancing to Stage 2 receive an Excellence Award (an Honorable Mention). | |
·Stage 2: Following target selectivity validation, the Organizing Committee will announce the wet-lab validation results for the 5 teams advancing to the Finals. |
3. Finals (Mid-October 2025):
The top five qualifying teams advancing to the Finals will be invited to deliver on-site oral presentations. Their presentations will be evaluated by the final judging panel, with each team’s scores announced live.
Based on the weighted total scores -comprising 70% from wet-lab validation results and 30% from oral presentation performance-the organizer will determine the winners of the first, second, and third Awards.
Important Notes:
Detailed defense requirements and scoring criteria will be published on the competition platform prior to the finals. Participants should monitor official announcements for updates.
(B) Large Molecule Track (Lactate Receptor-Specific Antibodies) Schedule:
1.Registration(From now - July 12 ,2025, 23:59:59 UTC/GMT+8)
Participants may register as individuals or in teams (all entrants must form a registered team). All required submissions must be completed within the specified timeframe. The organizing committee will announce the selected teams and the list of antibody molecule by July 25, 2025.
Required submissions from participating teams include:
Time | What to Submit | Requirements |
From now - July 12 ,2025, 23:59:59 UTC/GMT+8 | Proposal abstract | Participating teams must submit a proposed summary for their planned research using the official template. (Attachment 1. Template for Proposed Project Abstract.docx). The Organizing Committee will select maximum of 50 teams based on research competency (prior work in relevant fields), problem comprehension, implementation feasibility (methodological soundness) and innovation potential (novelty of approach). Please find detailed scoring rubric in Attachment 2. Abstract Scoring Criteria.pdf. |
Candidate molecules | Participating teams will compete using self-developed AI computational models, submitting a list of 50 molecules ranked by predicted score from highest to lowest. Based on this submitted prioritization, the Organizing Committee will select a minimum of 5 molecules per team for wet-lab validation.Please download template (Attachment 3. Preliminary Round Submission Template.docx) | |
The source code of the algorithm capable of reproducing molecule screening results | The Organizing Committee strongly encourages all participating teams to submit reproducible source code for molecular screening, promote open-source licensing and advocates for original algorithm development. Teams selected to advance in the competition must submit their source code. |
2.Preliminary Stage (July 25th –early October, 2025)
The organizers will conduct wet-lab validation and evaluation of submitted molecules (Attachment 6. Experimental Testing Procedure and Scoring Criteria for the Large Molecule Preliminary Round.pdf).
The Organizing Committee will announce the teams advancing to the Finals in mid-October 2025.
The Preliminary Round consists of the following three phases:
Time | Phase | Agenda |
July 25 –early October , 2025 | Screening | Based on dual-concentration antagonistic activity evaluation using GloSensor cAMP assay, the top 50 active molecules(ranked by antagonistic activity) will advance to the next phase. |
Evaluation | Following multi-concentration antagonistic activity evaluation and preliminary comparison using GloSensor cAMP assay, the top 10 ranked teams will advance. | |
Review | ·Stage 1: After multi-concentration antagonistic activity evaluation using HTRF cAMP assay, the top 5 ranked teams advance to Stage 2(and simultaneously qualify for the Finals). Teams entering the Review phase but not advancing to Stage 2 receive an Excellence Award (an Honorable Mention). | |
·Stage 2: Following target selectivity validation, the Organizing Committee will announce the wet-lab validation results for the 5 teams advancing to the Finals. |
3. Finals (Mid-October 2025):
The top five qualifying teams advancing to the Finals will be invited to deliver on-site oral presentations. Their presentations will be evaluated by the final judging panel, with each team’s scores announced live.
Based on the weighted total scores -comprising 70% from wet-lab validation results and 30% from oral presentation performance-the organizer will determine the winners of the first, second, and third Awards.
Important Notes:
Detailed defense requirements and scoring criteria will be published on the competition platform prior to the finals. Participants should monitor official announcements for updates.
VI. Competition Communication
If you have any questions about this competition, please feel free to contact us through the following channels:
Option 1: Add our official WeChat assistant
Scan the QR code to add the “Competition Assistant” account on WeChat.
After adding the account, you may send your inquiries directly via message.

Option 2: Contact us via email
Email: SICBC@yicai.com
To help us identify your message, please include “Competition Inquiry” in the email
subject line.
Important Notes:
1. Our inquiry service hours are Monday to Friday, 9:30 AM to 6:00 PM (working days only). Messages received during service hours will be processed and responded to as soon as possible.
Inquiries sent on non-working days (including public holidays), or outside service hours, will be handled on the next working day.
2. To ensure efficient communication and accurate information processing, please clearly indicate your full name, team name, and contact details (e.g., mobile number) when making an inquiry.
This information will be used solely by the Organizing Committee for internal identity verification and administrative purposes.
3. The above-mentioned WeChat account and email address are the official contact channels for this competition, and are exclusively operated and managed by Yicai Media Group.
Please remain vigilant and beware of counterfeit accounts or fraudulent impersonation.
VII. Organizers
Guidance: Science and Technology Commission of Shanghai Municipality
Host: Shanghai Center of Biomedicine Development
Co-hosts: Fudan University, East China Normal University, Shanghai Institute of Materia Medica.Chinese Academy of Sciences, Hengrui Pharma, Novoprotein, Shanghai SSCI Leading Private Equity Fund Management Co., Ltd., GUOTAI HAITONG SECURITIES Co., Ltd.,PHAIMUS,China Telecommunications Corporation, Shanghai Yicai Wanxiang Information Technology Co Ltd
Organizers: Shanghai Biopharma Service Company Limited,Shanghai Technological Service Platform for Biomedicine Industry
Co-organizer: Shanghai Titan Scientific Co.,LTD
VIII. Additional Rules
The following violations will result in disqualification by the Organizing Committee:
1. Fraudulent Registration: Submission of false information or failure to meet competition eligibility/team formation requirements.
2. Intellectual Property Violations: Submission of plagiarized work or infringement of third-party IP rights.
3. General Misconduct: Any other illegal or non-compliant activities identified during the competition period, whether through discovery or formal reporting.
4. Data Usage Restrictions: Competition-provided datasets are strictly authorized for model training purposes only. Commercial use of competition data is expressly prohibited.
5. IP Compliance: All intellectual property terms are governed by the Intellectual Property Annex (see bottom of page).
Final interpretation rights belong to the Organizing Committee.
| Documents | Action |
|---|---|
| 附件-知识产权说明 (Attachment. Intellectual Property Statement).zip |
Academic Committee of the Competition

Kaixian CHEN
Academician of the Chinese Academy of Sciences and Research Fellow at the Shanghai Institute of Materia Medica, Chinese Academy of Sciences

Xingming ZHAO
Professor at the Institute of Science and Technology for Brain-Inspired Intelligence, Fudan University

Haitao SONG
President of the Shanghai Artificial Intelligence Research Institute

Jiemin WENG
Dean of the School of Life Sciences, East China Normal University

Sheng WANG
Research Fellow at the Center for Excellence in Molecular Cell Science, Chinese Academy of Sciences

Huaxing ZHU
Chairman and General Manager of Novoprotein

Lianyi HAN
Head of the AIDD Department, Hengrui Pharma, Professor

Fang BAI
Professor at the School of Life Science and Technology, ShanghaiTech University
>>> Click to Access the Cloud Resource User Guide <<<
The competition organizers provide participating teams with access to cloud computing resources and related user guide. Resources are limited and will be distributed on a first-come, first-served basis.
[Pre-Competition Essentials]
I. Background of the Competition Topic
On February 4, 2025, a research team led by Researcher Weiqiang LU and Professor Mingyao LIU from the School of Life Sciences at East China Normal University published an article on Nature Immunology. The study was the first to reveal a novel mechanism by which the G protein–coupled receptor (GPCR) family member HCAR1 drives tumor immune evasion through lactate sensing. It also systematically demonstrated the potential application value of Lactate Receptor HCAR1 inhibitors in cancer immunotherapy.
Hydroxycarboxylic acid receptor 1 (HCAR1), a key member of the GPCR family, has emerged as a promising novel target for anti-cancer drug discovery. The development of potent and selective HCAR1-targeting molecules represents a highly valuable direction for innovative drug research. As a prototypical cell-surface receptor, HCAR1 can specifically recognize lactate molecules and mediate downstream signaling pathways. It plays a central role in regulating critical physiological processes such as energy metabolism and immune-inflammatory responses. Latest studies have shown that overactivation of HCAR1 signaling is closely associated with various pathological processes, including metabolic disorders and tumor progression. For instance, HCAR1 overactivation can enhance the recruitment of immunosuppressive cells, thereby inhibiting anti-tumor immune responses and driving malignant progression in cancers such as colorectal cancer. Additionally, abnormal activation of HCAR1 has been implicated in tumor-induced cachexia by promoting heightened metabolic activity in adipocytes.
At present, although the therapeutic potential of HCAR1 as a drug target is increasingly recognized, there remains a significant gap in the development of high-affinity, highly selective HCAR1-targeting drug molecules. The major obstacles hindering progress in this area include the technical challenges of high-throughput activity screening, the lack of detailed three-dimensional structural information, and interference from multiple homologous receptors.
This competition aims to address these challenges by leveraging the known protein sequence and structural data of HCAR1 to identify potential drug candidates through AI-driven screening of both small molecules (lactate receptor antagonists) and large molecules (HCAR1-specific antibodies). Candidate molecules will undergo wet-lab validation to assess their specificity and activity. By accelerating the discovery and translational development of HCAR1-targeted therapeutics, the competition seeks to advance the practical application of AI technologies in drug research and development.
II. Objective of the Competition Topic
Leveraging the known protein sequence and structural information of HCAR1 and related targets, the competition aims to discover highly active and selective HCAR1-targeting drug molecules by developing and optimizing AI-based computational methods, coupled with wet-lab validation. The ultimate goal is to promote the translational application of computational biology in innovative drug discovery.
III. Source of Drug Molecules
(I) Source of Compounds
1. Designated Compound Library:
The library comprises approximately 10 million compounds along with detailed information (a download link to the compound library will be provided: Computational Compound Library.zip). It includes around 20,000 compounds with known bioactivity, and approximately 10 million additional compounds drawn from a curated commercial selection and an extended compound collection.
(Library construction criteria: (1) Deduplication based on structure standardization and hash indexing methods; (2) Removal of unrecognizable molecules using the RDKIT algorithm; (3) Retention of compounds with molecular weights between 300 and 600 Da; and (4) Software compatibility testing.)
SMILES | MolWt | HBD | HBA | LogP | RotB | TPSA | Fsp3 | inchi | inchikey |
COC[C@H](CO)Nc1cccc(Cl)c1OC | 245.706 | 2 | 4 | 1.7678 | 6 | 50.72 | 0.454545 | 1S/C11H16ClNO3/c1-15-7-8(6-14)13-10-5-3-4-9(12)11(10)16-2/h3-5,8,13-14H,6-7H2,1-2H3/t8-/m0/s1 | BAOMYBXPPWJJFJ-QMMMGPOBSA-N |
2. Potential Sources of Antibody Sequences for Model Training:
The INDI database (http://research.naturalantibody.com) may be used by participating teams to perform high-throughput screening of candidate antibodies that bind to HCAR1, or to modify existing sequences derived from the database.
(II) Submission of Compounds by Participating Teams Encouraged: To incentivize the discovery of original innovative drugs, the competition accepts novel compound samples synthesized de novo by participating teams.
(III) Protein Sequence: The target protein sequence for this competition has been pre-specified.
(IV) Protein Structure: Participants may refer to the published HCAR1 structures (PDB IDs: 9J8Z, 9IZD) or perform homology modeling and other structure prediction methods.
IV. Submission of Entries
Submission Period: With effect from 23:59:59 on July 12, 2025
The competition requires each participating team to utilize a self-developed AI computational model. All submission materials must be uploaded via the “Submit Entry” page on the competition platform:
(I) Project Abstract: After registration, teams must submit a one-page abstract outlining their proposed approach (download abstract template: Attachment 1. Template for Proposed Project Abstract.docx; and scoring criteria: Attachment 2. Abstract Scoring Criteria.pdf).
(II) Submitted Molecules: Using their self-developed AI models, teams must submit 100 candidate molecules from each compound library. Molecules must be ranked in descending order based on predicted scores (Attachment 3. Preliminary Round Submission Template.docx).
(III) Source Code Model: The organizers encourage participating teams to submit reproducible source code models along with necessary introductory documents (e.g., model/code principles, code usage instruction). Submission of source code is mandatory for teams that advance to the final round.
Important Notes:
1. Small Molecule Compounds: For molecules submitted by participating teams that originate from the designated compound library, the organizers will be responsible for procurement or custom synthesis. If a compound is unavailable, has a long lead time, or is infeasible to synthesize, the next molecule on the team's submission list will be selected. For de novo synthesized compounds provided by teams, physical samples (10 mg) must be delivered to the organizers by participating teams before July 15, 2025. Teams must also submit corresponding characterization data, including structural formula, ¹H NMR, ¹³C NMR, high-resolution mass spectrometry (HRMS), and HPLC purity analysis.
2. Antibodies: Teams are only required to submit antibody sequences; no physical samples are needed. The organizers will handle all expression and activity validation. For standard consistency, antibody numbering must follow the IMGT (International ImMunoGeneTics) scheme, and the binding epitope of the antibody is restricted to the extracellular domain of the HCAR1 receptor.
3. Duplicate Submissions: If the same molecule is submitted by multiple teams, priority will be given to the team that submitted it first. The organizers will ensure that each team has at least 5-10 unique molecules accepted into the preliminary round.
4. Use of Open-Source Resources: The use of open-source data and code as a foundation is permitted, but submissions must clearly demonstrate original innovation. If source code is submitted, teams must explicitly cite the origin of any open-source data, code, or reference models used.
V. Scoring Criteria
(I) Scoring Criteria for Small Molecules:
Based on the project abstracts submitted by participating teams, the organizers will select no more than 100 teams to advance to the competition. Each team's final score will be calculated as a weighted sum of the preliminary round score (70%, based on wet-lab experimental results) and the final round score (30%, based on presentation performance). Final rankings will be determined based on these combined scores.
The scoring criteria for the preliminary round (Please dowload: Attachment 4. Experimental Testing Procedure and Scoring Criteria for the Small Molecule Preliminary Round.pdf, Attachment 5. Experimental Screening Workflow for Small Molecules.pdf) and the final round (Please dowload: Attachment 8. Final Round Scoring Criteria.pdf) are as follows:
1. Preliminary Round Scoring: All submitted molecules will undergo three stages of wet-lab evaluation: Screening, Primary Evaluation, and Secondary Evaluation. The evaluation criteria for each stage are as follows:
(1) Screening: Based on the prioritization of molecules submitted by participating teams, the organizers will select at least 10 molecules per team for high-throughput screening using the GloSensor cAMP assay, totaling approximately 1,000 molecules. The results will be ranked by antagonistic activity, and the top 100 molecules will advance to the competition.
(2) Primary Evaluation: The shortlisted molecules will undergo multi-concentration screening using the GloSensor cAMP assay. Molecules will be ranked by antagonistic activity, and molecules from the top 10 teams will proceed to the secondary evaluation. Priority will be given to teams with molecules among the top 10, with no more than three molecules per team entering the next stage. Additional teams will be added to ensure a total of 10 teams, following a differential completion principle, with only one molecule per team allowed.
(3) Secondary Evaluation:
Phase I (Antagonistic Activity Assessment): Molecules in this stage will be evaluated for their IC₅₀ values against HCAR1 using the HTRF cAMP assay. Based on the resulting scores, the top five teams (one molecule per team) will proceed to Phase II and qualify for the final round.
Phase II (Selectivity Assessment): The selected molecules will undergo further HTRF cAMP assays to measure IC₅₀ values against other GPCR family members, including HCAR2. These results will be scored according to detailed evaluation criteria, generating the final wet-lab performance score for each team.
2. Final Round Scoring
The five teams advancing to the final round will be invited to deliver on-site presentations. Each team will be scored by the Final Round Judging Panel, and the presentation scores will be announced on site.
(II) Scoring Criteria for Large Molecules:
Based on the project abstracts submitted by participating teams, the organizers will select no more than 50 teams to advance to the competition. Each team’s final score will be calculated as a weighted sum of the preliminary round score (70%, based on wet-lab experimental results) and the final round score (30%, based on presentation performance). Final rankings will be determined according to these combined scores.
The scoring criteria for the preliminary round (Please dowload: Attachment 6. Experimental Testing Procedure and Scoring Criteria for the Large Molecule Preliminary Round.pdf, Attachment 7. Experimental Screening Workflow for Large Molecules.pdf) and the final round (Please dowload: Attachment 8. Final Round Scoring Criteria.pdf) are as follows:
1. Preliminary Round Scoring: All submitted macromolecules will undergo three stages of wet-lab evaluation: Screening, Primary Evaluation, and Secondary Evaluation. The evaluation criteria for each stage are as follows:
(1) Screening: Based on the prioritization of molecules submitted by participating teams, the organizers will select no fewer than 5 molecules per team for initial screening. The organizers will first perform a preliminary expression level screening; only antibodies that meet the expression threshold will proceed to the subsequent high-throughput GloSensor cAMP assay. Based on the antagonistic activity results from this assay, the top 50 molecules will be selected to advance.
(2) Primary Evaluation: Antibodies that pass the initial screening will first undergo specific binding validation using flow cytometry to ensure selective binding to the HCAR1 receptor. Antibodies that pass this step will be further evaluated through multi-concentration screening using the GloSensor cAMP assay. Results will be ranked based on activity, and molecules from the top 10 teams will proceed to the secondary evaluation. Priority will be given to teams with molecules among the top 10 overall, with no more than three molecules per team entering the next stage. Additional teams will be added to ensure a total of 10 teams, following a differential completion principle, with only one molecule per team allowed.
(3) Secondary Evaluation:
Phase I (Antagonistic Activity Assessment): Molecules in this stage will be evaluated for their IC₅₀ values against HCAR1 using the HTRF cAMP assay. Based on the resulting scores, the top five teams (one molecule per team) will proceed to Phase II and qualify for the final round.
Phase II (Selectivity Assessment): The selected molecules will undergo further HTRF cAMP assays to measure IC₅₀ values against other GPCR family members, including HCAR2. These results will be scored according to detailed evaluation criteria, generating the final wet-lab performance score for each team.
2. Final Round Scoring: The five teams advancing to the final round will be invited to deliver on-site presentations. Each team will be scored by the Final Round Judging Panel, and the presentation scores will be announced on site.
| Documents | Action |
|---|---|
| 附件1 拟开展方案摘要模板 (Attachment 1. Template for Proposed Project Abstract).zip | |
| 附件2 摘要评分规则 (Attachment 2. Abstract Scoring Criteria).zip | |
| 附件3 初赛作品提交模板 (Attachment 3. Preliminary Round Submission Template).zip | |
| 附件4 小分子初赛实验测试流程与评分规则 (Attachment 4. Experimental Testing Procedure and Scoring Criteria for the Small Molecule Preliminary Round).zip | |
| 附件5 小分子实验筛选流程图 (Attachment 5. Experimental Screening Workflow for Small Molecules).zip | |
| 附件6 大分子初赛实验测试流程与评分规则 (Attachment 6. Experimental Testing Procedure and Scoring Criteria for the Large Molecule Preliminary Round).zip | |
| 附件7 大分子实验筛选流程图 (Attachment 7. Experimental Screening Workflow for Large Molecules).zip | |
| 附件8 决赛评分规则 (Attachment 8. Final Round Scoring Criteria).zip |
Awards and Benefits
The award ceremony is scheduled to be held during the International Biopharma Industry Week Shanghai 2025 (please refer to the official website of the competition for updates). The organizers will present awards to the winning teams and confer the following prizes and corresponding benefits. The award categories and associated benefits are as follows:
(I) Award Categories:
(1) Small Molecule Track (Lactate Receptor Antagonists):
1. First Prize (1 team):
CNY100,000 and an official award certificate.
2. Second Prize (2 teams):
CNY30,000 and an official award certificate per team.
3. Third Prize (2 teams):
CNY20,000 and an official award certificate per team.
4. Excellence Award (5 teams):
CNY10,000 and an official award certificate per team.
(2) Large Molecule Track (Lactate Receptor-Specific Antibodies):
1. First Prize (1 team):
CNY100,000 and an official award certificate.
2. Second Prize (2 teams):
CNY30,000 and an official award certificate per team.
3. Third Prize (2 teams):
CNY20,000 and an official award certificate per team.
4. Excellence Award (5 teams):
CNY10,000 and an official award certificate per team.
(II) Award Benefits for Recipients:
1. Preferential Recommendation. Award-winning projects will be given priority consideration for the “Computational Biology Special Program” under the “Shanghai Science and Technology Innovation Action Plan”, sponsored by the Science and Technology Commission of Shanghai Municipality.
2. Scenario-Based Collaboration. Promote market validation and real-world application of winning solutions through conferences and media exposure. Eligible winning enterprises will be supported to join Shanghai’s "Little Giant" SME Gradient Cultivation Program. Priority participation in drafting algorithmic standards and releasing research/consultancy reports for applied scenarios.
3. Commercialization Support. Priority access to domestic leading securities firms and venture capital funds, to facilitate the investment and accelerate commercialization of outstanding projects.
4. Academic Publication. The competition encourages and supports the publication of the competition results and related analyses in peer-reviewed academic journals.
Notes:
1. All prize amounts are pre-tax. Prizes will be disbursed to the winning teams after applicable taxes are withheld in accordance with relevant tax regulations.
2. If a single team submits multiple winning molecules, only the molecule with the highest-ranking award will be considered for the prize. Awards are non-cumulative. For example, a team that wins the First Prize will not receive an additional Excellence Award for other molecules it submitted.
3. In light of potential uncontrollable factors during the competition (such as variations in molecular activity or procurement timelines), the organizers reserve the right to adjust the competition rules during the competition according to the actual situation. The final interpretation of the competition rules rests solely with the organizers.
I. Competition Resources
Each participating team will be offered one cloud computing voucher valued at CNY2,000 (limited availability, distributed on a first-come, first-served basis).
Minimum Configuration Per Team:
Computing Resources: 6-core CPU | 30 GB RAM
GPU Resources: NVIDIA A10 ×1
Storage Resources: 100 GB
Usage Recommendation: Teams are advised to allocate and manage computing resources efficiently based on project needs.
Application Deadline: 23:59:59 on July 10, 2025
Usage Period: From now to 23:59:59 on July 12, 2025
Note:
The competition is team-based, and only team captains may request competition resources. Moreover,only one cloud computing voucher will be issued per team to a designated team member. Once team formation is confirmed, team members should not be changed.
II. Optional "Mirror Reference"
VNC | Minimal Image | Ubuntu:20.04 A pristine VNC image with Ubuntu desktop only
Jupyter | CUDA 12.1 | PyTorch 2.3.1 Preconfigured environment for model development
VS Code Web | PyTorch 2.3.1 | CUDA 12.1 Web-based VS Code environment for model development
III. Computing Resource Application Procedure
1. Application Procedure
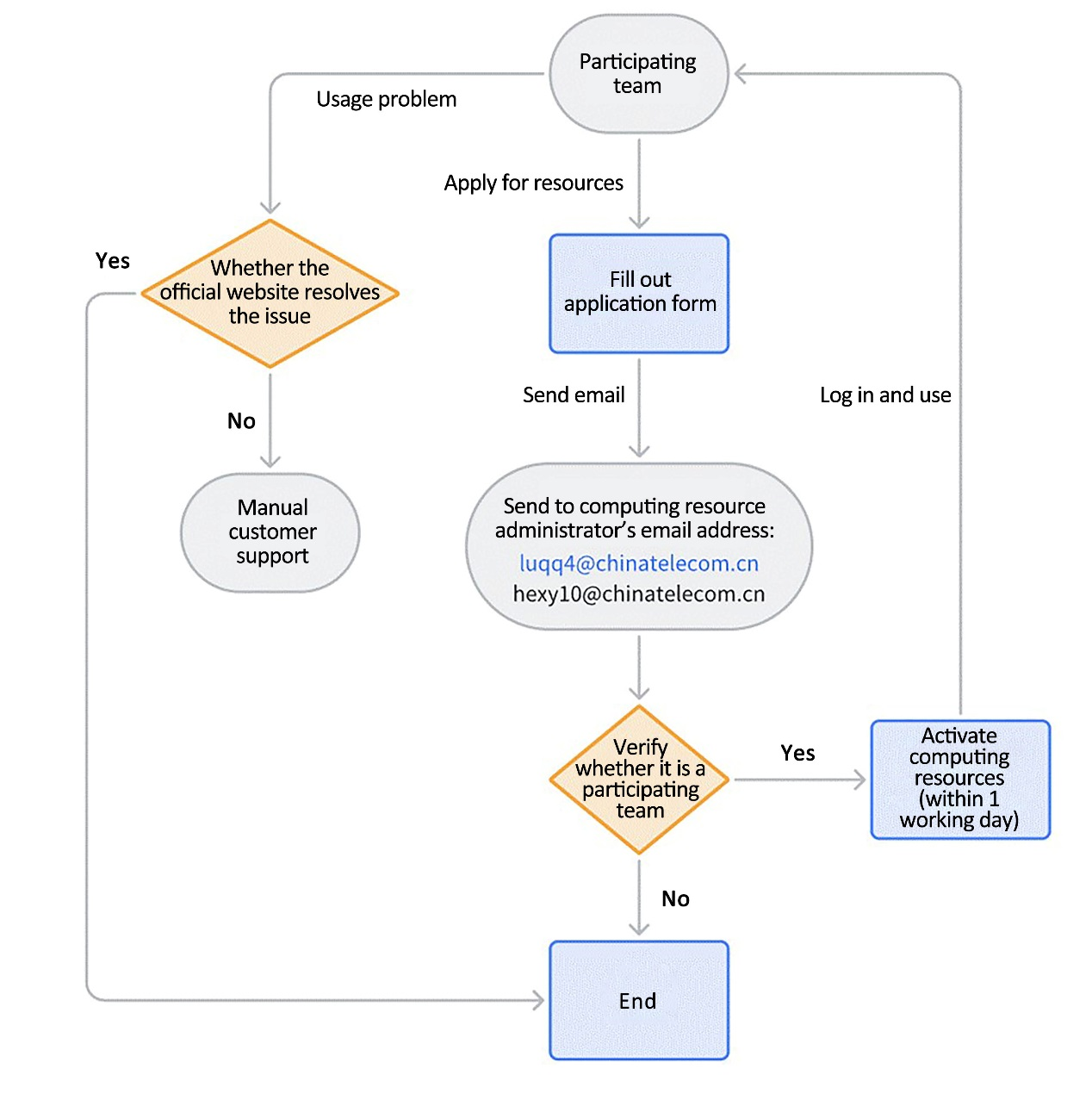
2. Example of Application Email
Subject: Application for Computing Resources—Computational Biology Competition 2025
Mail Address:
luqq4@chinatelecom.cn
hexy10@chinatelecom.cn
Email Body:
We are a participating team of the Computational Biology Competition 2025.
Team Name: [Insert Team Name]
We hereby apply for [Specify Resource Name] to support our research for the competition.
Applicant Information:
- Captain Name:
- Captain Mobile:
- Email:
(Note: Do not use an email address that has been registered for a eSurfing Cloud account.)
-Images: See available image references.
Please kindly approve our application.
Notes:
Minimum configuration: A10 specification. Instances will be provisioned based on current resource concurrency.
Incomplete submissions will receive verification requests from the Compute Resource Manager via email. Deployment confirmation will be provided upon completion.
IV. Relevant Materials
1. Access to the Computing Resource Platform: State Cloud Xirang – Research Assistant
https://www.ctyun.cn/products/batchcompute
2. User Manual for the Computing Platform:
https://www.ctyun.cn/document/10097674/%E7%94%A8%E6%88%B7%E6%8C%87%E5%8D%97
3. For inquiries, please contact the support personnel of the Computing Platform:
· Qingqing LU: 13817243557
· Xiaoyu HE: 19117234478
Create a team
Join the Team
| Team ID | Team Name | Track | Team Leader | Action | ||
|---|---|---|---|---|---|---|
| ${ v.id } | ${ v.name } | ${ v.racetrack } | ${ v.nickname } | ${ v.member_num } | Apply to join Fully staffed | Apply to join |
The application to join the team has been submitted and is waiting for the team leader's review
My team
Team member list(Limited to 5 people)
| Name | Phone Number | Role |
|---|---|---|
| ${ v.nickname } | ${ v.tel } | ${ v.is_leader ? 'Team Leader' : 'Team Member' } |
Team member application list
| Name | Phone Number | School | Major | Qualification | Action |
|---|---|---|---|---|---|
| ${ v.nickname } | ${ v.tel } | ${ v.school } | ${ v.major } | ${ v.qualification } |
*Upload works
Source code model
${ formWorks.racetrack === '大分子' ? 'Macromolecule' : 'Micromolecule' } Track
${ v.title }
${ formWorks.racetrack === '大分子' ? 'Macromolecule' : 'Micromolecule' } Track
${ v.title }
${ formWorks.racetrack === '大分子' ? 'Macromolecule' : 'Micromolecule' } Track
${ formWorks.racetrack === '大分子' ? 'Macromolecule' : 'Micromolecule' } Track

Scan the code to add WeChat assistant